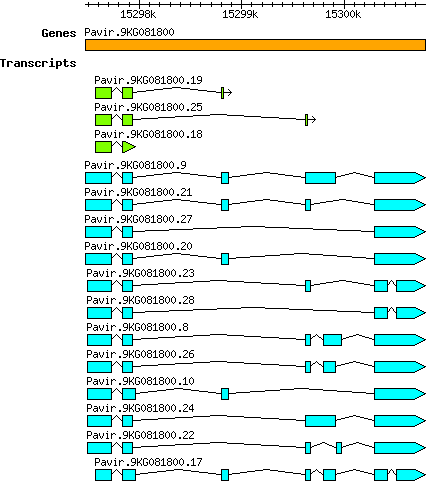
Pvirgatum Pavir.9KG081800
From GreeNC
Gene information
| Gene alias / name | Chrom. | Start | End | Assembly | Species | Details
|
|---|---|---|---|---|---|---|
| Pvirgatum_Pavir.9KG081800 [ Pavir.9KG081800] |
Chr09K | 15297474 | 15300794 | v5.1 | Panicum virgatum (Phytozome 13) | Coding? No |
Transcript features
This table shows how a lncRNA has been classified (high/low confidence) or whether it has been classified as pre-miRNA (go to FAQ for classification criteria). Besides, it displays the lncRNA length and sequence as well as information about the ORF, the coding potential calculated by the Coding Potential Calculator (CPC), and the folding energies calculated by RNAfold.
| Transcript alias / name | Confidence | Pre-miRNA | Length | Orthologous group | Sequence | Details
|
|---|---|---|---|---|---|---|
| Pvirgatum_Pavir.9KG081800.10 Pavir.9KG081800.10 |
Low | No | 930 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.230 Folding energies... MFEI: -36.312 AMFE: -0.674 GC content %: 53.871 | |
| Pvirgatum_Pavir.9KG081800.17 Pavir.9KG081800.17 |
Low | No | 931 | OG0021282 | Get ORF Coding potential... Type: noncoding Potential: -1.184 Folding energies... MFEI: -35.682 AMFE: -0.682 GC content %: 52.309 | |
| Pvirgatum_Pavir.9KG081800.18 Pavir.9KG081800.18 |
High | No | 285 | OG0282285 | Get ORF Coding potential... Type: noncoding Potential: -0.882 Folding energies... MFEI: -44.000 AMFE: -0.583 GC content %: 75.439 | |
| Pvirgatum_Pavir.9KG081800.19 Pavir.9KG081800.19 |
High | No | 273 | OG0282286 | Get ORF Coding potential... Type: noncoding Potential: -0.789 Folding energies... MFEI: -45.055 AMFE: -0.600 GC content %: 75.092 | |
| Pvirgatum_Pavir.9KG081800.20 Pavir.9KG081800.20 |
Low | No | 910 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.088 Folding energies... MFEI: -35.681 AMFE: -0.664 GC content %: 53.736 | |
| Pvirgatum_Pavir.9KG081800.21 Pavir.9KG081800.21 |
Low | No | 962 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.101 Folding energies... MFEI: -34.865 AMFE: -0.655 GC content %: 53.222 | |
| Pvirgatum_Pavir.9KG081800.22 Pavir.9KG081800.22 |
Low | No | 923 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.117 Folding energies... MFEI: -36.316 AMFE: -0.681 GC content %: 53.304 | |
| Pvirgatum_Pavir.9KG081800.23 Pavir.9KG081800.23 |
Low | No | 786 | OG0021282 | Get ORF Coding potential... Type: noncoding Potential: -1.078 Folding energies... MFEI: -38.244 AMFE: -0.672 GC content %: 56.870 | |
| Pvirgatum_Pavir.9KG081800.24 Pavir.9KG081800.24 |
Low | No | 1114 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.158 Folding energies... MFEI: -33.654 AMFE: -0.672 GC content %: 50.090 | |
| Pvirgatum_Pavir.9KG081800.25 Pavir.9KG081800.25 |
High | No | 270 | OG0282287 | Get ORF Coding potential... Type: noncoding Potential: -0.812 Folding energies... MFEI: -45.667 AMFE: -0.604 GC content %: 75.556 | |
| Pvirgatum_Pavir.9KG081800.26 Pavir.9KG081800.26 |
Low | No | 1000 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.135 Folding energies... MFEI: -35.490 AMFE: -0.682 GC content %: 52.000 | |
| Pvirgatum_Pavir.9KG081800.27 Pavir.9KG081800.27 |
Low | No | 841 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.145 Folding energies... MFEI: -35.791 AMFE: -0.656 GC content %: 54.578 | |
| Pvirgatum_Pavir.9KG081800.28 Pavir.9KG081800.28 |
Low | No | 734 | OG0021282 | Get ORF Coding potential... Type: noncoding Potential: -1.115 Folding energies... MFEI: -38.801 AMFE: -0.672 GC content %: 57.766 | |
| Pvirgatum_Pavir.9KG081800.8 Pavir.9KG081800.8 |
Low | No | 1057 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.147 Folding energies... MFEI: -34.844 AMFE: -0.678 GC content %: 51.372 | |
| Pvirgatum_Pavir.9KG081800.9 Pavir.9KG081800.9 |
Low | No | 1204 | OG0002705 | Get ORF Coding potential... Type: noncoding Potential: -1.148 Folding energies... MFEI: -32.508 AMFE: -0.655 GC content %: 49.668 |
Matches to external databases
| Transcript alias | Database | Version | Hit | Evalue |
|---|---|---|---|---|
| Pvirgatum_Pavir.9KG081800.26 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.20 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.27 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.21 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.28 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.22 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.8 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.23 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.10 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia | |
| Pvirgatum_Pavir.9KG081800.9 | RepBase (RepeatMasker) | 2014-01-31 | DNA/PIF-Harbinger; LTR/Copia |
... further results
Gene models