Gene
Csativus Cucsa.350950
From GreeNC
Gene information
| Gene alias / name | Chrom. | Start | End | Source | Assembly | Species | Details
|
|---|---|---|---|---|---|---|---|
| Csativus_Cucsa.350950 Cucsa.350950 |
scaffold03504 | 367002 | 370587 | Phytozome | v1.0 | Cucumis_sativus | Coding? No |
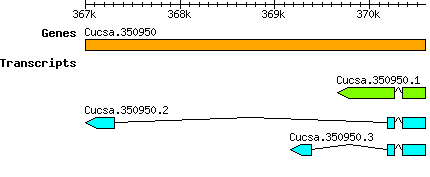
Transcript features
This table shows how a lncRNA has been classified (high/low confidence) or whether it has been classified as pre-miRNA (go to FAQ for classification criteria). Besides, it displays the lncRNA length and sequence as well as information about the ORF, the coding potential calculated by the Coding Potential Calculator (CPC), and the folding energies calculated by RNAfold.
| Transcript alias / name | Confidence | Pre-miRNA | Length | Sequence | Details
|
|---|---|---|---|---|---|
| Csativus_Cucsa.350950.1 Cucsa.350950.1 PAC:16979931 |
High | No | 851 | Get ORF Coding potential... Type: noncoding Potential: -0.532 Folding energies... MFEI: -25.817 AMFE: -0.684 GC content %: 37.720 | |
| Csativus_Cucsa.350950.2 Cucsa.350950.2 PAC:16979932 |
Low | No | 634 | Get ORF Coding potential... Type: noncoding Potential: -0.469 Folding energies... MFEI: -26.404 AMFE: -0.622 GC content %: 42.429 | |
| Csativus_Cucsa.350950.3 Cucsa.350950.3 PAC:16979933 |
Low | No | 543 | Get ORF Coding potential... Type: noncoding Potential: -0.407 Folding energies... MFEI: -25.506 AMFE: -0.630 GC content %: 40.516 |
Matches to external databases
| Transcript alias | Database | Version | Hit | Evalue |
|---|---|---|---|---|
| Csativus_Cucsa.350950.2 | SwissProt | 2015-06-01 | Q2M2S5 | 6.0e-4 |
| Csativus_Cucsa.350950.3 | SwissProt | 2015-06-01 | Q2M2S5 | 4.0e-4 |
Gene models